|
12/28/2023 0 Comments Sequencher contig assembly imverted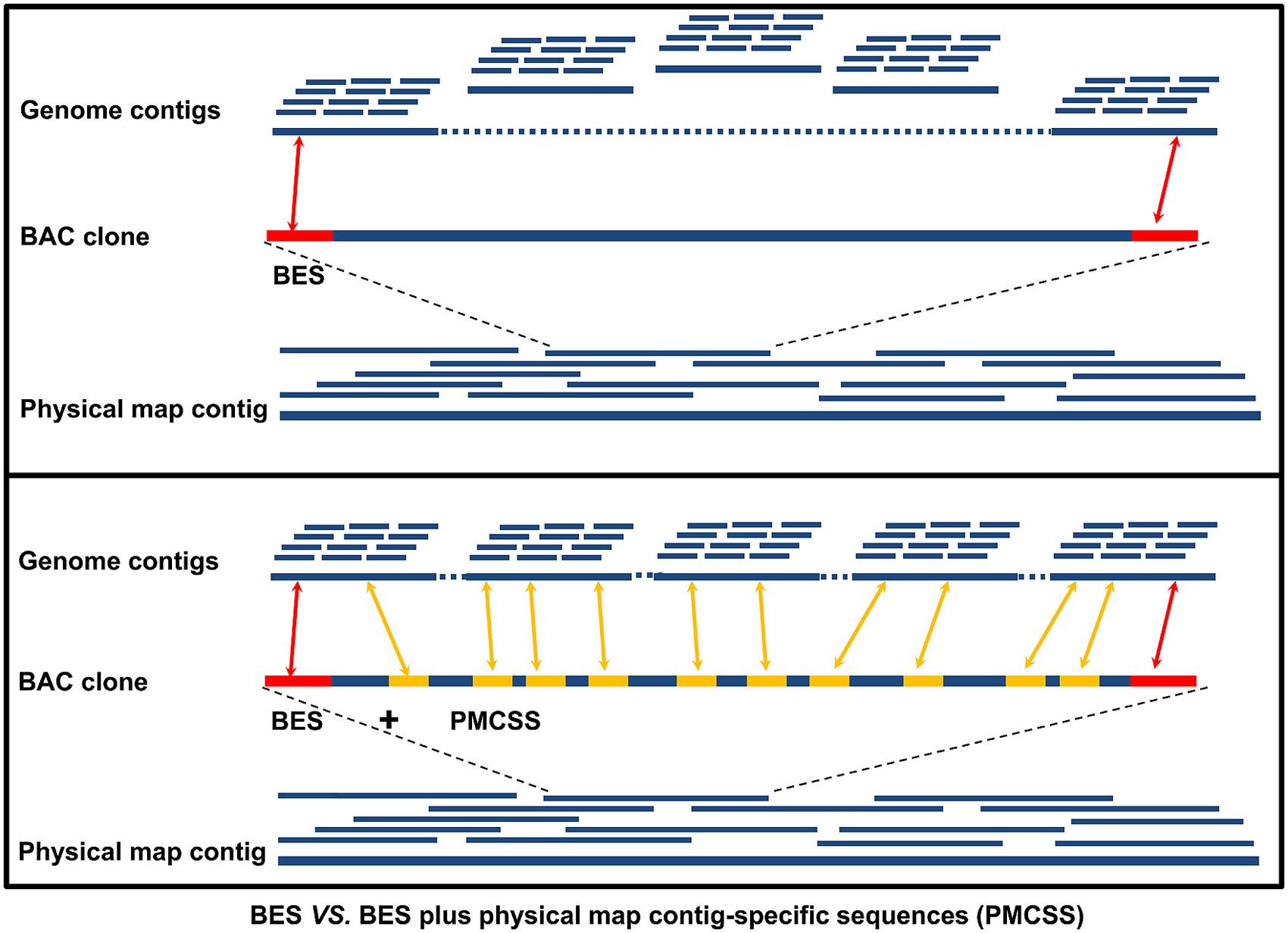
We hope this software will be useful and helpful for those without programming experience but with a large set of samples to analyze. Third, ambiguous nucleotides or gaps and the four junctional regions between the IRs and SSC/LSC in the chloroplast genome sequences were further confirmed by PCR amplification and Sanger sequencing with specific primers ( Dong et al., 2013 ). This program could greatly save time for the people. The selected contigs from chloroplast genomes were further assembled using Sequencher 5.4.5. The Classification function will make some preprocessing for the sequence files, and the Assembly function will generate a SeqMan script to consecutively assemble the contigs for the classified files. contig assembly, and custom reporting that is superior in nature to those. There are two modules available in autoSeqMan, and. Accuracy is greater than 99 using Phred 20 bi-directional sequence traces. Here, we present a software called to automatedly assemble contigs with program SeqMan. While, with increasing of data size, larger set of samples as well as more loci sequenced make the contig assembling very tricky, majorly because a lot of manual operations are needed by using SeqMan. SeqMan (in package DNASTAR) is an excellent program with an easy-to-use graphical user interface (GUI) to assemble the Sanger sequences into contigs. 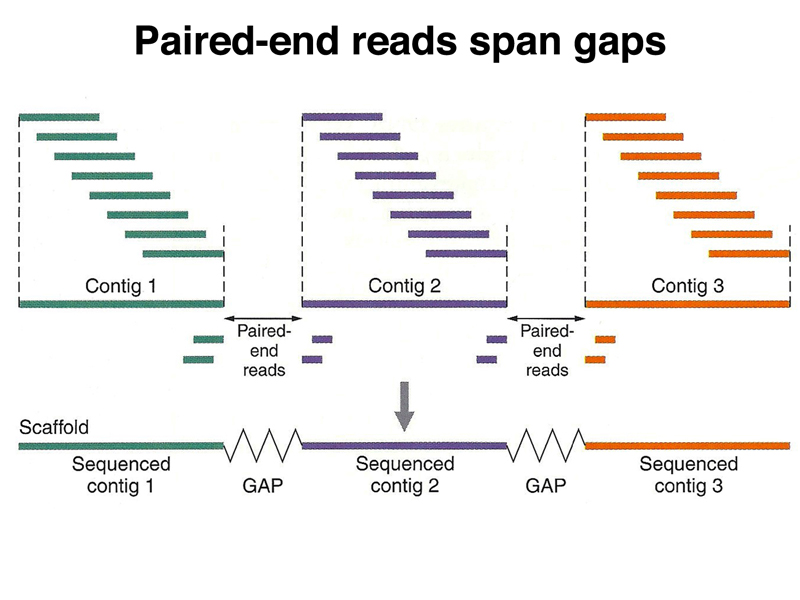
Mitochondrial contigs were manually connected into larger contigs using Sequencher 5.2 (Gene Codes Corp., Ann Arbor, MI, USA) based on consistent paired-end reads visualized in Consed v26. With the wide application of DNA barcoding technology, more and more DNA sequences are generated through Sanger sequencing platform. Genome assemblies and read-pair mapping patterns were visually inspected using Consed v26 (Gordon & Green, 2013). Sequencher from Gene Codes of Ann Arbor, Mich., specializes in contig assembly, but includes vector/transposon screening and heterozygote identification for. Compare contigs: Align contigs with MUSCLE or Clustal, and go back to the chromatograms with a simple mouse click. Unless removed by trimming, any of these artifacts will distort your sequence assembly and downstream sequence analysis. However, with increasing data size, larger sample sets and more sequenced loci make contig assemble complicated due to the considerable number of manual operations required to run SeqMan. AutoSeqMan Batch Assembling Contigs for Sanger Sequences Introns and primer sequence frequently flank the sequence of amplified exons. SeqMan (in the LaserGene package) is an excellent program with an easy-to-use graphical user interface (GUI) employed to assemble Sanger sequences into contigs.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
